Topologically associating domain

A topologically associating domain (TAD) is a self-interacting genomic region, meaning that DNA sequences within a TAD physically interact with each other more frequently than with sequences outside the TAD.[1] The median size of a TAD in mouse cells is 880 kb, and they have similar sizes in non-mammalian species.[2] Boundaries at both side of these domains are conserved between different mammalian cell types and even across species[2] and are highly enriched with CCCTC-binding factor (CTCF) and cohesin.[1] In addition, some types of genes (such as transfer RNA genes and housekeeping genes) appear near TAD boundaries more often than would be expected by chance.[3][4]
The functions of TADs are not fully understood and are still a matter of debate. Most of the studies indicate TADs regulate gene expression by limiting the enhancer-promoter interaction to each TAD;[5] however, a recent study uncouples TAD organization and gene expression.[6] Disruption of TAD boundaries are found to be associated with wide range of diseases such as cancer,[7][8][9] variety of limb malformations such as synpolydactyly, Cooks syndrome, and F-syndrome,[10] and number of brain disorders like Hypoplastic corpus callosum and Adult-onset demyelinating leukodystrophy.[10]
The mechanisms underlying TAD formation are also complex and not yet fully elucidated, though a number of protein complexes and DNA elements are associated with TAD boundaries. However, the handcuff model and the loop extrusion model describe the TAD formation by the aid of CTCF and cohesin proteins.[11] Furthermore, it has been proposed that the stiffness of TAD boundaries itself could cause the domain insulation and TAD formation.[11]
Discovery and diversity
[edit]TADs are defined as regions whose DNA sequences preferentially contact each other. They were discovered in 2012 using chromosome conformation capture techniques including Hi-C.[3][12][4] They have been shown to be present in multiple species,[13] including fruit flies (Drosophila),[14] mouse,[3] plants, fungi and human[4] genomes. In bacteria, they are referred to as Chromosomal Interacting Domains (CIDs).[13]
Analytical tools and databases
[edit]TAD locations are defined by applying an algorithm to Hi-C data. For example, TADs are often called according to the so-called "directionality index".[4] The directionality index is calculated for individual 40kb bins, by collecting the reads that fall in the bin, and observing whether their paired reads map upstream or downstream of the bin (read pairs are required to span no more than 2Mb). A positive value indicates that more read pairs lie downstream than upstream, and a negative value indicates the reverse. Mathematically, the directionality index is a signed chi-square statistic.
The development of specialized genome browsers and visualization tools[15] such as Juicebox,[16] HiGlass[17]/HiPiler,[18] The 3D Genome Browser,[19] 3DIV,[20] 3D-GNOME,[21] and TADKB[22] have enabled us to visualize the TAD organization of regions of interest in different cell types.
Mechanisms of formation
[edit]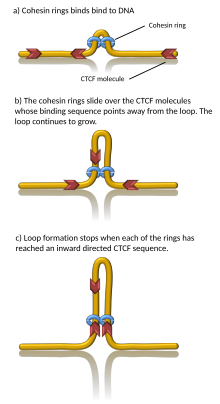
A number of proteins are known to be associated with TAD formation including the protein CTCF and the protein complex cohesin.[1] It is also unknown what components are required at TAD boundaries; however, in mammalian cells, it has been shown that these boundary regions have comparatively high levels of CTCF binding. In addition, some types of genes (such as transfer RNA genes and housekeeping genes) appear near TAD boundaries more often than would be expected by chance.[3][4]
Computer simulations have shown that chromatin loop extrusion driven by cohesin motors can generate TADs.[23][24] In the loop extrusion model, cohesin binds chromatin, pulls it in, and extrudes chromatin to progressively grow a loop. Chromatin on both sides of the cohesin complex is extruded until cohesin encounters a chromatin-bound CTCF protein, typically located at the boundary of a TAD. In this way, TAD boundaries can be brought together as the anchors of a chromatin loop.[25] Indeed, in vitro, cohesin has been observed to processively extrude DNA loops in an ATP-dependent manner[26][27][28] and stall at CTCF.[29][30] However, some in vitro data indicates that the observed loops may be artifacts.[31][32] Importantly, since cohesins can dynamically unbind from chromatin, this model suggests that TADs (and associated chromatin loops) are dynamic, transient structures,[23] in agreement with in vivo observations.[33][34][35][36]
Other mechanisms for TAD formation have been suggested. For example, some simulations suggest that transcription-generated supercoiling can relocalize cohesin to TAD boundaries[37][38] or that passively diffusing cohesin “slip links”[39][40] can generate TADs.
Properties
[edit]Conservation
[edit]TADs have been reported to be relatively constant between different cell types (in stem cells and blood cells, for example), and even between species in specific cases.[4][41][42][43]
Relationship with promoter-enhancer contacts
[edit]The majority of observed interactions between promoters and enhancers do not cross TAD boundaries. Removing a TAD boundary (for example, using CRISPR to delete the relevant region of the genome) can allow new promoter-enhancer contacts to form. This can affect gene expression nearby - such misregulation has been shown to cause limb malformations (e.g. polydactyly) in humans and mice.[42]
Computer simulations have shown that transcription-induced supercoiling of chromatin fibres can explain how TADs are formed and how they can assure very efficient interactions between enhancers and their cognate promoters located in the same TAD.[37]
Relationship with other structural features of the genome
[edit]Replication timing domains have been shown to be associated with TADs as their boundary is co localized with the boundaries of TADs that are located at either sides of compartments.[44] Insulated neighborhoods, DNA loops formed by CTCF/cohesin-bound regions, are proposed to functionally underlie TADs.[45]
Role in disease
[edit]Disruption of TAD boundaries can affect the expression of nearby genes, and this can cause disease.[46]
For example, genomic structural variants that disrupt TAD boundaries have been reported to cause developmental disorders such as human limb malformations.[47][48][49] Additionally, several studies have provided evidence that the disruption or rearrangement of TAD boundaries can provide growth advantages to certain cancers, such as T-cell acute lymphoblastic leukemia (T-ALL),[50] gliomas,[51] and lung cancer.[52]
Lamina-associated domains
[edit]
Lamina-associated domains (LADs) are parts of the chromatin that heavily interact with the lamina, a network-like structure at the inner membrane of the nucleus.[53] LADs consist mostly of transcriptionally silent chromatin, being enriched with trimethylated Lys27 on histone H3, (i.e. H3K27me3); which is a common posttranslational histone modification of heterochromatin.[54] LADs have CTCF-binding sites at their periphery.[53]
See also
[edit]References
[edit]- ^ a b c Pombo A, Dillon N (April 2015). "Three-dimensional genome architecture: players and mechanisms". Nature Reviews. Molecular Cell Biology. 16 (4): 245–257. doi:10.1038/nrm3965. PMID 25757416. S2CID 6713103.
- ^ a b Yu M, Ren B (October 2017). "The Three-Dimensional Organization of Mammalian Genomes". Annual Review of Cell and Developmental Biology. 33: 265–289. doi:10.1146/annurev-cellbio-100616-060531. PMC 5837811. PMID 28783961.
- ^ a b c d Nora EP, Lajoie BR, Schulz EG, Giorgetti L, Okamoto I, Servant N, et al. (April 2012). "Spatial partitioning of the regulatory landscape of the X-inactivation centre". Nature. 485 (7398): 381–385. Bibcode:2012Natur.485..381N. doi:10.1038/nature11049. PMC 3555144. PMID 22495304.
- ^ a b c d e f Dixon JR, Selvaraj S, Yue F, Kim A, Li Y, Shen Y, et al. (April 2012). "Topological domains in mammalian genomes identified by analysis of chromatin interactions". Nature. 485 (7398): 376–380. Bibcode:2012Natur.485..376D. doi:10.1038/nature11082. PMC 3356448. PMID 22495300.
- ^ Krijger PH, de Laat W (December 2016). "Regulation of disease-associated gene expression in the 3D genome". Nature Reviews. Molecular Cell Biology. 17 (12): 771–782. doi:10.1038/nrm.2016.138. PMID 27826147. S2CID 11484886.
- ^ Ghavi-Helm Y; Jankowski A; Meiers S; Viales RR; Korbel JO; Furlong EE (August 2019). "Highly rearranged chromosomes reveal uncoupling between genome topology and gene expression". Nature Genetics. 51 (8): 1272–1282. doi:10.1038/s41588-019-0462-3. PMC 7116017. PMID 31308546.
- ^ Corces MR, Corces VG (February 2016). "The three-dimensional cancer genome". Current Opinion in Genetics & Development. 36: 1–7. doi:10.1016/j.gde.2016.01.002. PMC 4880523. PMID 26855137.
- ^ Valton AL; Dekker J (February 2016). "TAD disruption as oncogenic driver". Current Opinion in Genetics & Development. 36: 34–40. doi:10.1016/j.gde.2016.03.008. PMC 4880504. PMID 27111891.
- ^ Achinger-Kawecka J, Clark SJ (January 2017). "Disruption of the 3D cancer genome blueprint". Epigenomics. 9 (1): 47–55. doi:10.2217/epi-2016-0111. PMID 27936932.
- ^ a b Spielmann M, Lupiáñez DG, Mundlos S (July 2018). "Structural variation in the 3D genome". Nature Reviews. Genetics. 19 (7): 453–467. doi:10.1038/s41576-018-0007-0. hdl:21.11116/0000-0003-610A-5. PMID 29692413. S2CID 22325904.
- ^ a b Dixon JR, Gorkin DU, Ren B (June 2016). "Chromatin Domains: The Unit of Chromosome Organization". Molecular Cell. 62 (5): 668–680. doi:10.1016/j.molcel.2016.05.018. PMC 5371509. PMID 27259200.
- ^ de Laat W, Duboule D (October 2013). "Topology of mammalian developmental enhancers and their regulatory landscapes". Nature. 502 (7472): 499–506. Bibcode:2013Natur.502..499D. doi:10.1038/nature12753. PMID 24153303. S2CID 4468533.
- ^ a b Szabo Q, Bantignies F, Cavalli G (April 2019). "Principles of genome folding into topologically associating domains". Science Advances. 5 (4): eaaw1668. Bibcode:2019SciA....5.1668S. doi:10.1126/sciadv.aaw1668. PMC 6457944. PMID 30989119.
- ^ Sexton T, Yaffe E, Kenigsberg E, Bantignies F, Leblanc B, Hoichman M, et al. (February 2012). "Three-dimensional folding and functional organization principles of the Drosophila genome". Cell. 148 (3): 458–472. doi:10.1016/j.cell.2012.01.010. PMID 22265598.
- ^ Ing-Simmons E, Vaquerizas JM (September 2019). "Visualising three-dimensional genome organisation in two dimensions". Development. 146 (19): 99–101. doi:10.1242/dev.177162. PMID 31558569.
- ^ Durand NC; Robinson JT; Shamim MS; Machol I; Mesirov JP; Lander ES; Aiden EL (July 2016). "Juicebox Provides a Visualization System for Hi-C Contact Maps with Unlimited Zoom". Cell Systems. 3 (1): 99–101. doi:10.1016/j.cels.2015.07.012. PMC 5596920. PMID 27467250.
- ^ Kerpedjiev P, Abdennur N, Lekschas F, McCallum C, Dinkla K, Strobelt H, et al. (August 2018). "HiGlass: web-based visual exploration and analysis of genome interaction maps". Genome Biology. 19 (1): 125. doi:10.1186/s13059-018-1486-1. PMC 6109259. PMID 30143029.
- ^ Lekschas F, Bach B, Kerpedjiev P, Gehlenborg N, Pfister H (January 2018). "HiPiler: Visual Exploration of Large Genome Interaction Matrices with Interactive Small Multiples". IEEE Transactions on Visualization and Computer Graphics. 24 (1): 522–531. doi:10.1109/TVCG.2017.2745978. PMC 6038708. PMID 28866592.
- ^ Wang Y, Song F, Zhang B, Zhang L, Xu J, Kuang D, et al. (October 2018). "The 3D Genome Browser: a web-based browser for visualizing 3D genome organization and long-range chromatin interactions". Genome Biology. 19 (1): 151. doi:10.1186/s13059-018-1519-9. PMC 6172833. PMID 30286773.
- ^ Yang D, Jang I, Choi J, Kim MS, Lee AJ, Kim H, et al. (January 2018). "3DIV: A 3D-genome Interaction Viewer and database". Nucleic Acids Research. 46 (D1): D52–D57. doi:10.1093/nar/gkx1017. PMC 5753379. PMID 29106613.
- ^ Szalaj P, Michalski PJ, Wróblewski P, Tang Z, Kadlof M, Mazzocco G, et al. (July 2016). "3D-GNOME: an integrated web service for structural modeling of the 3D genome". Nucleic Acids Research. 44 (W1): W288–W293. doi:10.1093/nar/gkw437. PMC 4987952. PMID 27185892.
- ^ Liu, T., Porter, J., Zhao, C. et al. TADKB: Family classification and a knowledge base of topologically associating domains. BMC Genomics 20, 217 (2019). https://doi.org/10.1186/s12864-019-5551-2
- ^ a b Fudenberg G, Imakaev M, Lu C, Goloborodko A, Abdennur N, Mirny LA (May 2016). "Formation of Chromosomal Domains by Loop Extrusion". Cell Reports. 15 (9): 2038–2049. doi:10.1016/j.celrep.2016.04.085. PMC 4889513. PMID 27210764.
- ^ Sanborn AL; Rao SS; Huang SC; Durand NC; Huntley MH; Jewett AI; et al. (November 2015). "Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes". Proceedings of the National Academy of Sciences of the United States of America. 112 (47): E6456–E6465. Bibcode:2015PNAS..112E6456S. doi:10.1073/pnas.1518552112. PMC 4664323. PMID 26499245.
- ^ Yatskevich S; Rhodes J; Nasmyth K (December 2019). "Organization of Chromosomal DNA by SMC Complexes". Annual Review of Genetics. 53 (1): 445–482. doi:10.1146/annurev-genet-112618-043633. PMID 31577909. S2CID 203653572.
- ^ Golfier S, Quail T, Kimura H, Brugués J (May 2020). Dekker, Struhl K, Mirny LA, Musacchio A, Marko JF (eds.). "Cohesin and condensin extrude DNA loops in a cell cycle-dependent manner". eLife. 9: e53885. doi:10.7554/eLife.53885. PMC 7316503. PMID 32396063.
- ^ Davidson IF, Bauer B, Goetz D, Tang W, Wutz G, Peters JM (December 2019). "DNA loop extrusion by human cohesin". Science. 366 (6471): 1338–1345. Bibcode:2019Sci...366.1338D. doi:10.1126/science.aaz3418. PMID 31753851. S2CID 208228309.
- ^ Kim Y, Shi Z, Zhang H, Finkelstein IJ, Yu H (December 2019). "Human cohesin compacts DNA by loop extrusion". Science. 366 (6471): 1345–1349. Bibcode:2019Sci...366.1345K. doi:10.1126/science.aaz4475. PMC 7387118. PMID 31780627.
- ^ Davidson IF, Barth R, Zaczek M, van der Torre J, Tang W, Nagasaka K, et al. (2022-09-09). "CTCF is a DNA-tension-dependent barrier to cohesin-mediated DNA loop extrusion". bioRxiv 10.1101/2022.09.08.507093.
- ^ Zhang H, Shi Z, Banigan EJ, Kim Y, Yu H, Bai X, Finkelstein IJ (2022-10-07). "CTCF and R-loops are boundaries of cohesin-mediated DNA looping". bioRxiv 10.1101/2022.09.15.508177.
- ^ Man, Zhou (September 2022). "DNA sliding and loop formation by E. coli SMC complex: MukBEF". Biochemistry and Biophysics Reports. 31: 101297. doi:10.1016/j.bbrep.2022.101297. PMC 9234588. PMID 35770038.
- ^ Ryu JK, Bouchoux C, Liu HW, Kim E, Minamino M, de Groot R, Katan AJ, Bonato A, Marenduzzo D, Michieletto D, Uhlmann F (February 2021). "Bridging-induced phase separation induced by cohesin SMC protein complexes". Science Advances. 7 (7): eabe5905. Bibcode:2021SciA....7.5905R. doi:10.1126/sciadv.abe5905. PMC 7875533. PMID 33568486.
- ^ Gabriele M, Brandão HB, Grosse-Holz S, Jha A, Dailey GM, Cattoglio C, et al. (April 2022). "Dynamics of CTCF- and cohesin-mediated chromatin looping revealed by live-cell imaging". Science. 376 (6592): 496–501. Bibcode:2022Sci...376..496G. doi:10.1126/science.abn6583. PMC 9069445. PMID 35420890.
- ^ Beckwith KS, Ødegård-Fougner Ø, Morero NR, Barton C, Schueder F, Tang W, et al. (2022-05-02). "Visualization of loop extrusion by DNA nanoscale tracing in single human cells". bioRxiv 10.1101/2021.04.12.439407.
- ^ Mach P, Kos PI, Zhan Y, Cramard J, Gaudin S, Tünnermann J, et al. (2022-03-03). "Live-cell imaging and physical modeling reveal control of chromosome folding dynamics by cohesin and CTCF". bioRxiv 10.1101/2022.03.03.482826.
- ^ Flyamer IM, Gassler J, Imakaev M, Brandão HB, Ulianov SV, Abdennur N, et al. (April 2017). "Single-nucleus Hi-C reveals unique chromatin reorganization at oocyte-to-zygote transition". Nature. 544 (7648): 110–114. Bibcode:2017Natur.544..110F. doi:10.1038/nature21711. PMC 5639698. PMID 28355183.
- ^ a b Racko D, Benedetti F, Dorier J, Stasiak A (January 2019). "Are TADs supercoiled?". Nucleic Acids Research. 47 (2): 521–532. doi:10.1093/nar/gky1091. PMC 6344874. PMID 30395328.
- ^ Racko D, Benedetti F, Dorier J, Stasiak A (February 2018). "Transcription-induced supercoiling as the driving force of chromatin loop extrusion during formation of TADs in interphase chromosomes". Nucleic Acids Research. 46 (4): 1648–1660. doi:10.1093/nar/gkx1123. PMC 5829651. PMID 29140466.
- ^ Brackley CA, Johnson J, Michieletto D, Morozov AN, Nicodemi M, Cook PR, Marenduzzo D (September 2017). "Nonequilibrium Chromosome Looping via Molecular Slip Links". Physical Review Letters. 119 (13): 138101. arXiv:1612.07256. Bibcode:2017PhRvL.119m8101B. doi:10.1103/PhysRevLett.119.138101. PMID 29341686. S2CID 14706723.
- ^ Yamamoto T, Schiessel H (September 2017). "Osmotic mechanism of the loop extrusion process". Physical Review E. 96 (3–1): 030402. Bibcode:2017PhRvE..96c0402Y. doi:10.1103/PhysRevE.96.030402. hdl:1887/58394. PMID 29346962.
- ^ Vietri Rudan M, Barrington C, Henderson S, Ernst C, Odom DT, Tanay A, Hadjur S (March 2015). "Comparative Hi-C reveals that CTCF underlies evolution of chromosomal domain architecture". Cell Reports. 10 (8): 1297–1309. doi:10.1016/j.celrep.2015.02.004. PMC 4542312. PMID 25732821.
- ^ a b Jost D, Vaillant C, Meister P (February 2017). "Coupling 1D modifications and 3D nuclear organization: data, models and function". Current Opinion in Cell Biology. 44: 20–27. doi:10.1016/j.ceb.2016.12.001. PMID 28040646.
- ^ Yang Y, Zhang Y, Ren B, Dixon JR, Ma J (June 2019). "Comparing 3D Genome Organization in Multiple Species Using Phylo-HMRF". Cell Systems. 8 (6): 494–505.e14. doi:10.1016/j.cels.2019.05.011. PMC 6706282. PMID 31229558.
- ^ Marchal C, Sima J, Gilbert DM (December 2019). "Control of DNA replication timing in the 3D genome". Nature Reviews. Molecular Cell Biology. 20 (12): 721–737. doi:10.1038/s41580-019-0162-y. PMID 31477886. S2CID 201714312.
- ^ Ji X, Dadon DB, Powell BE, Fan ZP, Borges-Rivera D, Shachar S, et al. (February 2016). "3D Chromosome Regulatory Landscape of Human Pluripotent Cells". Cell Stem Cell. 18 (2): 262–275. doi:10.1016/j.stem.2015.11.007. PMC 4848748. PMID 26686465.
- ^ Lupiáñez DG, Spielmann M, Mundlos S (April 2016). "Breaking TADs: How Alterations of Chromatin Domains Result in Disease". Trends in Genetics. 32 (4): 225–237. doi:10.1016/j.tig.2016.01.003. hdl:11858/00-001M-0000-002E-1D1D-D. PMID 26862051.
- ^ Lupiáñez DG, Kraft K, Heinrich V, Krawitz P, Brancati F, Klopocki E, et al. (May 2015). "Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions". Cell. 161 (5): 1012–1025. doi:10.1016/j.cell.2015.04.004. PMC 4791538. PMID 25959774.
- ^ Angier N (2017-01-09). "A Family's Shared Defect Sheds Light on the Human Genome". The New York Times.
- ^ Franke M, Ibrahim DM, Andrey G, Schwarzer W, Heinrich V, Schöpflin R, et al. (October 2016). "Formation of new chromatin domains determines pathogenicity of genomic duplications". Nature. 538 (7624): 265–269. Bibcode:2016Natur.538..265F. doi:10.1038/nature19800. hdl:11858/00-001M-0000-002C-010A-3. PMID 27706140. S2CID 4463482.
- ^ Hnisz D, Weintraub AS, Day DS, Valton AL, Bak RO, Li CH, et al. (March 2016). "Activation of proto-oncogenes by disruption of chromosome neighborhoods". Science. 351 (6280): 1454–1458. Bibcode:2016Sci...351.1454H. doi:10.1126/science.aad9024. PMC 4884612. PMID 26940867.
- ^ Flavahan WA, Drier Y, Liau BB, Gillespie SM, Venteicher AS, Stemmer-Rachamimov AO, et al. (January 2016). "Insulator dysfunction and oncogene activation in IDH mutant gliomas". Nature. 529 (7584): 110–114. Bibcode:2016Natur.529..110F. doi:10.1038/nature16490. PMC 4831574. PMID 26700815.
- ^ Weischenfeldt J, Dubash T, Drainas AP, Mardin BR, Chen Y, Stütz AM, et al. (January 2017). "Pan-cancer analysis of somatic copy-number alterations implicates IRS4 and IGF2 in enhancer hijacking". Nature Genetics. 49 (1): 65–74. doi:10.1038/ng.3722. PMC 5791882. PMID 27869826.
- ^ a b Gonzalez-Sandoval A; Gasser SM (August 2016). "On TADs and LADs: Spatial Control Over Gene Expression". Trends in Genetics. 32 (8): 485–495. doi:10.1016/j.tig.2016.05.004. PMID 27312344.
- ^ Li M, Liu GH, Izpisua Belmonte JC (July 2012). "Navigating the epigenetic landscape of pluripotent stem cells". Nature Reviews. Molecular Cell Biology. 13 (8): 524–535. doi:10.1038/nrm3393. PMID 22820889. S2CID 22524502.
